Featured
A Data Biosphere for Biomedical Research
By Benedict Paten
UCSC Genomics Institute
October 18, 2017 — Santa Cruz, CA
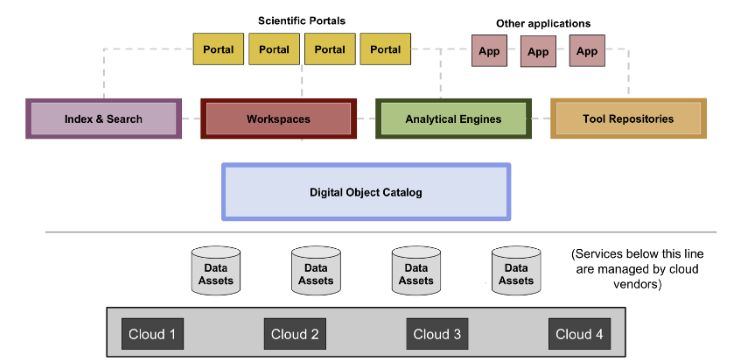
(Image above: The diagram illustrates a proposed architecture for a Data Biosphere. More details here.)
We, the authors listed below, are privileged to be part of the growing global community bringing data and life science together. Our groups have been working together in overlapping combinations during the past two years to drive the creation of data commons to support flagship scientific initiatives. This document lays out our evolving vision for the next steps in that journey. Our hope is that others will join the effort to build momentum for an open, compatible, and secure approach to data within the larger research community. We welcome your feedback, and look forward to continuing this journey together.
Josh Denny (Vanderbilt), David Glazer (Verily Life Sciences), Robert L. Grossman (University of Chicago), Benedict Paten (University of California at Santa Cruz), Anthony Philippakis (Broad Institute)
Introduction
Data is playing an ever greater role in the life sciences. Thirty years ago, the data that most biomedical researchers needed resided in their lab notebooks. Today, research projects are often informed by vast stores of data — from technologies such as genome sequencing, gene-expression analysis, imaging, and high-throughput chemical screens — generated by individual laboratories and community-wide projects around the world.
These massive datasets, posted in repositories across the Internet, are a boon to experimentalists seeking to interpret their own results, as well as to the growing cadre of computational biologists looking for patterns that only emerge by looking at the big picture. But, they pose huge challenges. It can be difficult to find and download the datasets, interpret their formats, and perform computations combining diverse information. Moreover, as datasets grow in scale, the practice of downloading data is becoming impractical in terms of cost (storing multiple copies of large datasets is wasteful), accessibility (few researchers have the necessary computational infrastructure) and security (many research laboratories lack state-of-the-art security and access control).
The obvious solution is cloud-based data storage and computation designed for biomedical research. But, how should it be designed?
One approach is to create a closed, monolithic data platform. This model is conceptually simple and has precedents in the world of technology. But, it’s the wrong answer for the life sciences. Closed systems are contrary to the spirit of the scientific enterprise. And, monolithic systems won’t provide the flexibility to accommodate the wide range of scientific needs and the diverse regulatory requirements of different jurisdictions.
Instead, we propose the idea of creating a vibrant ecosystem, which we call the “Data Biosphere.” It contains modular and interoperable components that can be assembled into diverse data environments. In the following we outline how the biomedical community can work together to rapidly achieve this goal.
Data Biosphere: Principles
The Data Biosphere should be based on four governing principles. It should be: (1) modular, composed of functional components with well-specified interfaces; (2) community-driven, created by many groups to foster a diversity of ideas; (3) open, developed under open-source licenses that enable extensibility and reuse, with users able to add custom, proprietary modules as needed; and (4) standards-based, consistent with standards developed by coalitions such as the Global Alliance for Genomics and Health (GA4GH). As in biological ecosystems, the health of the Data Biosphere will be measured by both its activity and its diversity.
Continue reading here: https://medium.com/@benedictpaten/a-data-biosphere-for-biomedical-research-d212bbfae95d
###
Tagged Genomics Institute, UC Santa Cruz








